Abstract
The development of paired appendages was a key innovation during evolution and facilitated the aquatic to terrestrial transition of vertebrates. Largely derived from the lateral plate mesoderm (LPM), one hypothesis for the evolution of paired fins invokes derivation from unpaired median fins via a pair of lateral fin folds located between pectoral and pelvic fin territories1. Whilst unpaired and paired fins exhibit similar structural and molecular characteristics, no definitive evidence exists for paired lateral fin folds in larvae or adults of any extant or extinct species. As unpaired fin core components are regarded as exclusively derived from paraxial mesoderm, any transition presumes both co-option of a fin developmental programme to the LPM and bilateral duplication2. Here, we identify that the larval zebrafish unpaired pre-anal fin fold (PAFF) is derived from the LPM and thus may represent a developmental intermediate between median and paired fins. We trace the contribution of LPM to the PAFF in both cyclostomes and gnathostomes, supporting the notion that this is an ancient trait of vertebrates. Finally, we observe that the PAFF can be bifurcated by increasing bone morphogenetic protein signalling, generating LPM-derived paired fin folds. Our work provides evidence that lateral fin folds may have existed as embryonic anlage for elaboration to paired fins.
Similar content being viewed by others
Main
Two alternate hypotheses have been proposed to explain the evolutionary origin of vertebrate paired appendages (fins and limbs). Derivation from posterior gill arches was posited by Gegenbaur3, whilst a number of anatomists later invoked a rival theory, the lateral fin fold hypothesis. This proposed that paired fins derived (either phylogenetically or ontogenetically) from longitudinal bilateral fin folds that were then subdivided4,5. Whilst recent molecular studies have provided some evidence in support of each hypothesis2,6,7, there remains significant criticism of the lack of substantiation in the fossil record or in embryology1,8. Certain stem vertebrates, including anaspid-related fossils, show evidence of lateral fin folds; however, these fin folds mostly consist of soft tissue with only sporadic skeletal elements and are thus poorly preserved. This has led to conflicting interpretations9,10,11,12,13. The developmental programme for paired fins has been postulated to have been first assembled in median fins, which appear in the fossil record before the origin of paired fins2. A number of studies have traced the cellular origin of median fins in lamprey, catshark, zebrafish and Xenopus2,14,15. All median fins, both larval and adult, assessed so far have shown derivation from the paraxial mesoderm (PM), whilst paired fins are known to be derived from the lateral plate mesoderm (LPM). Thus, the median fin programme was most likely transferred to the LPM from the PM2, possibly before the formation of hypothesized lateral fin folds. How or when this transition occurred is unclear. As only a subset of median fins in zebrafish has been assayed, we expanded the characterization of the composition and origin of median fins in zebrafish to determine if PM derivation was an invariant characteristic.
As with most surveyed jawed vertebrates (gnathostomes), larval zebrafish possess two median unpaired fin folds. A caudal fin fold (or major lobe) runs continuously from the dorsal midline around the caudal end of the larva and then ventrally to the anus (Fig. 1a). A pre-anal fin fold (PAFF; or minor lobe) runs along the underside of the yolk sac extension, immediately anterior to the anus16 (Fig. 1a). The median fin folds are resorbed during metamorphosis, and the caudal fin fold is replaced by three separate PM-derived median adult fins14,17: the caudal fin, the dorsal fin and the anal fin. No adult median fin replaces the PAFF, which is a developmentally transient structure16. The mesenchyme of the caudal fin fold can be labelled by the photoconvertible Kaede green-to-red fluorescent protein expressed under control of the tbx16l promoter, confirming that this median fin is derived from the PM18. In contrast, we consistently failed to observe any Kaede labelling in the PAFF (Fig. 1b and Extended Data Fig. 1a). This labelling difference is not due to de novo activity of the tbx16l promoter specifically in the caudal median fin fold, as we photoconverted Kaede in anterior somites above the PAFF at 24 h post-fertilization (hpf) and traced the photoconverted Kaede to the mesenchyme of the dorsal portion of the caudal fin fold; again, we never found labelled cells in the PAFF (n = 11) (Extended Data Fig. 1b–d). The absence of PAFF labelling by a PM-lineage trace is not due to absence of mesenchyme cells, as we observed an abundance of these cells by Nomarski optics at 3 days post-fertilization (dpf) and in the enhancer trap line Et(krt4:EGFP)sqet37 (ET37), which labels all fin mesenchyme cells (Extended Data Fig. 1e,f). The ET37 line further allowed us to visualize the morphology of the mesenchyme cells, which showed a distally polarized stellate shape, indistinguishable from the mesenchyme of all other fins (Fig. 1c–e and Extended Data Fig. 1f–h). PAFF mesenchymal cells expressed known differentiation markers of fin mesenchyme, including fibulin1 (fbln1) and integrin beta 3b (itgb3b), and were weakly positive for lyve1b reporter activity in the lyve1b:DsRed2 transgenic line, which also expresses in the post-anal ventral fin mesenchyme19 (Fig. 1f–h and Extended Data Fig. 1i). Hence, despite their divergent developmental origin, PAFF mesenchyme has a similar morphology and expression profile to fin mesenchyme of the major median fin lobe.
a, Larval 4 dpf zebrafish possess a median PAFF (yellow arrowhead) in addition to the caudal median fin fold (cyan arrowhead). b, Confocal image of a 3 dpf Tg(tbx16l:GAL4-VP16); Tg(UAS:Kaede) embryo with PM labelled by Kaede showing PM-derived mesenchyme in the caudal median fin fold (cyan outline) but not the PAFF (yellow outline). c–e, Confocal images of pre-anal (c), ventral caudal (d) and pectoral (e) fins of the ET37 Enhancer Trap transgenic line indicating that PAFF contains morphologically comparable mesenchyme (indicated by arrowheads) to other larval fin folds. f, In situ hybridization of the fin mesenchyme marker fbln1 in both PAFF (yellow arrowhead) and caudal fin fold (cyan arrowhead) at 3 dpf. g,h, DsRed expression in mesenchyme of both pre-anal (g) and caudal (h) fin folds of the 5 dpf Tg(-5.2lyve1b:DsRed) transgenic line. i,j, Immunostaining for collagen II in 8 dpf frf mutants (j) shows loss of fibril organization compared with WT (i). Scale bars, 200 µm (a); 100 µm (b); 20 µm (c,e); 50 µm (f,g,j).
Few functions of fin mesenchyme cells are defined. The zebrafish frilly fin (frf) mutants display ruffling of the caudal larval fin fold due to mutations in bone morphogenetic protein 1a (bmp1a), which is expressed in fin mesenchyme20, including in the PAFF (Extended Data Fig. 1j). The overall morphology of the PAFF also displays ruffling in frf mutants (Fig. 1i,j). Further, immunostaining of collagen II revealed ordered parallel collagen II fibres in all median fins of the wild type (WT), while these fibres were disorganized in both the caudal median fin folds and the PAFF of frf mutants (Fig. 1i,j), indicating that Bmp1a in PAFF mesenchyme is also required for maturation of collagen. Collectively, these results indicate that mesenchymal cells of the PAFF show functional overlap with those of the caudal median fin fold yet have a distinct developmental origin.
The transcription factor Hand2 is expressed in several LPM progenitors, including cardiac, pharyngeal, mesothelial and pectoral fin progenitors21,22. Upon examining the TgBAC(hand2:EGFP) transgenic line that accurately recapitulates endogenous hand2 gene expression23, we observed the expected enhanced green fluorescent protein (eGFP) expression in larval pectoral fins at 2–3 dpf (Fig. 2a,b). Although we observed no TgBAC(hand2:EGFP) expression in the major unpaired larval fin lobe, the PAFF mesenchyme was strongly eGFP positive, suggesting an LPM origin (Fig. 2a,b and Extended Data Fig. 2a,b). Additionally, in situ hybridization at 3 and 5 dpf indicated that the PAFF mesenchyme, but no other median fin fold, expressed hand2 (Fig. 2c,d and Extended Data Fig. 2c,d). If the LPM uniquely contributes to the PAFF, then we might expect that the PAFF, but no other median fin fold, would be affected in an LPM mutant. The zebrafish hand2 mutant hands off (hans6) exhibits defects in LPM derivatives, including the heart, pectoral fins and mesothelium21,22. Consistent with an LPM origin, the fin height of PAFFs in hans6 mutant larvae was significantly reduced compared with WT at 3 and 5 dpf, whilst the ventral section of the major median fin lobe was unaffected (Extended Data Fig. 3a,b,g,h,m). We observed similar results with hand2 antisense morpholino (MO)-based knockdown (Extended Data Fig. 3c–f,n). hand2 knockdown in the ET37 line severely reduced the number of eGFP-positive mesenchymal cells in the PAFF at 3 and 5 dpf compared with WT, whilst the mesenchyme of the ventral caudal median fin fold was unaffected (Extended Data Fig. 3i–l,o). Transplantation of hand2 MO; ET37 mutant cells into WT suggested that the loss of Hand2 reduced clone size and altered morphology and migration of mesenchyme cells autonomously (Extended Data Fig. 4). Together, these observations are consistent with an LPM origin of PAFF mesenchyme.
a,b, Confocal images of Tg(hand2:EGFP) embryos at 2 dpf (a) and 3 dpf (b) showing eGFP labelling of the mesenchyme of the pectoral (green outline and arrowheads) and PAFFs (yellow outline and arrowheads, and magnified in inset) but not the caudal fin fold (cyan outline). c,d, In situ hybridization of hand2 at 3 dpf shows fin expression of hand2 only in the PAFF (yellow arrowheads (c), and higher magnification with Nomarski optics indicates expression in the mesenchyme (d). e, Schematic of the LPM lineage tracing transgenes. f, Lineage tracing of LPM using transgenics depicted in (e) following 4-OHT treatment and heat shock before imaging shows that PAFF mesenchyme is derived from the LPM (yellow arrowhead and magnified in inset). g,h, Ventral (g) and lateral (h) confocal images of the drl:H2B-Dendra2 transgenic line at the 10-somite stage (10 ss) (g) and 48 hpf (h) following ultra-violet laser photoconversion in the region of the LPM outlined in (g). h, Photoconverted PAFF mesenchyme is indicated by yellow arrowheads. Scale bars, 100 µm (a,c,f); 50 µm (a (inset),f (inset),g,h); 200 µm (b); 20 µm (d).
To confirm that this expression of hand2 indicates an LPM origin of the PAFF rather than de novo expression, we permanently labelled the LPM in a mosaic fashion using a Tg(drl:creERT2; hsp70l:Switch) transgenic combination24 (Fig. 2e). During gastrulation and early somitogenesis, drl:creERT2 is expressed in LPM-primed mesoderm with increasing selectivity to the LPM, suitable for lineage labelling of this mesodermal lineage using 4-OH-tamoxifen (4-OHT) induction and loxP reporters24. Consistent with an LPM origin of paired appendages, we first documented that this LPM lineage tracing approach labels cells of the paired fins, in particular the larval pectoral fins and pelvic fins. We found lineage labelling in the endoskeletal disc and mesenchyme of 4.5 dpf pectoral fins and also, the intraray fibroblasts and Zns5-positive osteoblasts of the adult pelvic fins (Extended Data Fig. 5a–d). In addition, we observed LPM lineage labelling of the PAFF mesenchyme at 3 and 5 dpf, whilst the labelling in the caudal fin fold was limited to myeloid cells, which are also LPM derived (Fig. 2f and Extended Data Fig. 5e–g). These data further support the contribution of the LPM to the PAFF mesenchyme but not to other median fin folds. We next used drl-driven expression of the Dendra2 green-to-red photoconvertible fluorophore to localize the precise LPM origin of PAFF mesenchyme. During early to mid-somitogenesis (8–10-somite stage (ss)), drl:H2B-Dendra2 is expressed in the nuclei of the LPM (Extended Data Fig. 6a,b). Photoconversion of posterior-most LPM at this stage, using ultra-violet laser illumination, resulted in selective Dendra2-red labelling of this field of LPM (Fig. 2g and Extended Data Fig. 6c,d). These cells were then traced to the migrating PAFF mesenchyme at 40 and 48 hpf (n = 4) (Fig. 2h and Extended Data Fig. 6e–j,q–t), whilst unconverted control cells did not show any Dendra2-red PAFF labelling (Extended Data Fig. 6k–p,u–x). This confirmed PAFF mesenchyme originates from the LPM, with significant contribution from the posterior-most LPM at early segmentation stages.
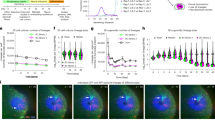
To investigate whether an LPM-seeded median PAFF is a zebrafish apomorphy or if it is a more ancestral feature shared with other vertebrates, we examined the larvae of other taxa for the presence of a PAFF and used hand2 in situ hybridization as a proxy marker of an LPM origin of constituent mesenchymal cells. Medaka (Oryzias latipes) larvae have a markedly smaller PAFF that possesses only a sparse number of mesenchymal cells as detected by Nomarski imaging (Fig. 3a,b). These mesenchymal cells have cell extensions projecting distally and were mostly proximal in the fin where the interstitial space is thickest (Fig. 3b). Accordingly, medaka hand2 was expressed proximally in a punctate pattern in PAFFs, consistent with mesenchymal expression, and was not expressed in other median fin folds (Fig. 3c and Extended Data Fig. 7a–c). To confirm this, we expressed eGFP in early medaka LPM by injection of a -6.35drl:EGFP construct. We saw expression in mesenchymal cells in the PAFF transiently and also, in stable transgenic embryos (eight PAFF mesenchyme cells seen in four stage 36 Tg(-6.35drl:EGFP) transgenic medaka embryos) (Extended Data Fig. 7d,e). In addition, we used 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate (DiI) lipophilic dye labelling to label the posterior LPM territory at stage 20 (four-somite stage). Consistent with our positional mapping of PAFF-seeding mesenchyme in zebrafish, posterior LPM injection of DiI consistently labelled mesenchyme in the medaka PAFF (total of 27 cells in nine of nine labelled embryos) (Extended Data Fig. 7f–i). A basal actinopterygian, the American paddlefish (Polyodon spathula), also forms a transient larval PAFF8; we identified the expression of P. spathula hand2 in PAFF mesenchymal cells between stages 36 and 39, but hand2 expression was notably absent from the caudal median fin fold (Fig. 3d,e and Extended Data Fig. 7j–l). PAFF hand2 mesenchyme expression persisted until PAFF regression (around stages 45 and 46). From stage 39, we observed hand2 expression in the core of the nascent pelvic fins (Extended Data Fig. 7k,l). We conclude that a larval PAFF with an LPM mesenchyme is ancestral for actinopterygians. To establish if an LPM contribution to median fins occurred before the origin of paired fins, we examined the larvae of a jawless vertebrate, the sea lamprey (Petromyzon marinus). Lampreys also possess a small transient larval PAFF25, within which strong, specific expression of the lamprey hand2 orthologue, HandA, was detected (Fig. 3f,g), although there was evidence of fainter HandA expression in the dorsal anterior fin. Finally, both the embryos of the sarcopterygian lungfish26 and amphibian tadpoles (e.g., Xenopus laevis27, Xenopus tropicalis and axolotls15) also possess a small transient PAFF (Fig. 3h). Fluorescence in situ hybridization demonstrated individual hand2-positive mesenchymal cells invading this fin in X. tropicalis but not the major median fin lobe at stage 42 (Fig. 3h–j). We confirmed this LPM contribution by injecting -6.35drl:EGFP into X. laevis embryos. X. laevis has a larger PAFF than X. tropicalis, and we observed extensive transient eGFP labelling in the PAFF compared with the caudal fin fold (Extended Data Fig. 7m–o). Thus, we show an LPM origin for PAFF mesenchyme in the larvae of representatives of major extant vertebrate linages: cyclostomes, actinopterygians and sarcopterygians.
a,b, Nomarski images of 9 dpf medaka showing PAFF (a) with dispersed mesenchymal cells (yellow arrowheads) (b). c, In situ hybridization of medaka at stage 39 showing hand2 expression in pre-anal fin mesenchyme (yellow arrowheads). d,e, In situ hybridization of hand2 in stage 36 paddlefish embryos shown laterally (d) or in transverse section (e). hand2-positive PAFF fin mesenchyme is indicated by yellow arrowheads. f,g, HandA in situ hybridization of stage E29 lamprey (P. marinus) embryos shown laterally (f) or in transverse section (g). Lamprey show strong expression of HandA in a fin anterior to the anus (yellow arrowhead in f) corresponding to cells in the interior of the fin (g). The section location of (g) is indicated by the dashed line in (f). h–j, Chromogenic (h) or fluorescent (i,j) in situ hybridization of hand2 in stage 42 X. tropicalis embryos shown laterally (h) or in transverse section (i,j). Fluorescent image (j) is overlayed on the Nomarski image (i). The small PAFF in Xenopus contains sparse hand2-positive fin mesenchyme (yellow arrowheads). St., stage. Scale bars, 100 µm (a,d,h); 10 µm (b); 20 µm (c); 50 µm (e–g,i).
Given the LPM origin of the median PAFF, we considered that it may have importance in the transition from PM-derived median fins to LPM-derived paired fins. We tested if this LPM median fin fold could be duplicated into paired LPM-derived lateral fin folds. Zebrafish larvae mutant for the bone morphogenetic protein (BMP) antagonist chordin develop ventral caudal fin fold duplications, although whether the PAFF is duplicated in these mutants has not been reported28. Injection of a low dose of chrd MO into zebrafish recapitulated various ventralized phenotypes, which included individuals with duplicated or multiple PAFFs (Fig. 4a,b,f and Extended Data Fig. 8a,b). We visualized these PAFF phenotypes by chrd MO injection into the ET37 transgenic line, which demonstrated that each fin fold of duplicated PAFFs contained mesenchyme (Fig. 4c,d). Lineage tracing using the Tg(drl:creERT2; hsp70l:Switch) transgenic combination injected with chrd MO demonstrated that the mesenchyme cells of the duplicated PAFFs are indeed derived from the LPM (Fig. 4e). By generating chrd morphants in the Tg(hand2:EGFP) line and then imaging by light-sheet microscopy, we generated three-dimensional reconstructions documenting duplicated PAFFs along the yolk extension, all of which harboured eGFP-positive mesenchymal cells, further demonstrating that the mesenchyme of these duplicate fin folds was LPM derived (Fig. 4g–i, Extended Data Fig. 8c–j and Supplementary Video 1). Of note is the generation of multiple parallel fin folds in individual embryos following chrd MO injection (Fig. 4f). These observations indicate that bifurcation of the LPM-seeded PAFF readily arises from modulating ventral BMP signalling, such as by reducing the dose of the dorsal inhibitor Chordin.
a,b, Lateral (a) and ventral (b) Nomarski images of PAFFs of 5 dpf (a) and 4 dpf (b) embryos injected with MO targeting chordin, resulting in duplication of PAFF (yellow arrowheads). c,d, Confocal micrographs, imaged ventrally, of PAFFs in 4 dpf ET37 transgenic larvae uninjected (c) or injected with a chrd MO (d). PAFFs are indicated with yellow lines. d, Note the duplicated anal openings following chrd MO injection. e, Ventral confocal image of chrd MO duplicated PAFFs of the transgenic line shown above. Hydroxytamoxifen treatment at the 12-somite stage and heat shock before imaging show that mesenchyme of duplicated PAFFs is derived from the LPM (yellow arrowheads). f–i, Light-sheet (f) and confocal (g–i) images of Tg(hand2:EGFP) larvae at 8 dpf (f) and 6 dpf (g–i). Orthogonal display through the x–z plane (f,g) shows that multiple PAFFs can form and that duplicated PAFFs contain eGFP-positive mesenchyme (g–i). Lateral views of left (h) and right (i) duplicated PAFFs of sample in g are given. j,k, Lateral low- (j) and high-power (k) Nomarski images of the PAFF of the Ranchu goldfish at 6 dpf. The multiple PAFFs (j) and individual mesenchyme cells (k) are indicated by yellow arrowheads. Duplicated caudal fin folds are indicated by cyan arrowheads (j). l–n, Lateral (l,m) and transverse (n) views of pre-anal (l,n) and caudal (m) fin folds of 7 dpf Ranchu larvae stained by in situ hybridization for hand2, where hand2-positive PAFF mesenchyme is indicated by yellow arrowheads. Absence of hand2 in the caudal fin fold is indicated by the cyan arrowhead (m). Scale bars, 100 µm (a,e,m); 20 µm (b,f–h,n); 50 µm (d,k); 200 µm (j).
To determine if the generation of multiple PAFFs could have evolved spontaneously as part of natural variation, we examined the twin-tail goldfish strain, Ranchu, which has been shown to display bifurcated fin folds including PAFFs due to a loss-of-function mutation in a chordin paralogue (chdAE127X)29,30,31. The bifurcated PAFFs and caudal fin folds of larval Ranchu appeared similar to those of zebrafish larvae injected with low doses of chrd MO (Fig. 4j and Extended Data Fig. 8k,l). These PAFFs were all populated by mesenchymal cells (Fig. 4k). Using hand2 in situ hybridization as a proxy of LPM lineage to infer the origin of the Ranchu PAFF mesenchyme, we observed that the core of both duplicated PAFFs expressed hand2, whilst the caudal fin fold did not (Fig. 4l–n and Extended Data Fig. 8m,n). This suggests that, as in the zebrafish PAFF, the bifurcated Ranchu PAFFs are also LPM derived. Notably, in addition to those displaying simple PAFF duplication, some individuals had three or more parallel PAFFs (Fig. 4j and Extended Data Fig. 8n). Thus, we conclude that LPM-derived paired fin folds that can be readily generated in zebrafish larvae have also arisen spontaneously in twin-tail goldfish; thus, they represent a viable morphological innovation. Notably, duplicated LPM-derived PAFFs were situated in ventrolateral locations between the future pectoral and pelvic fin domains, overlying the proposed competence stripes of appendage formation in gnathostomes32.
Combining several models and techniques, we here have identified a larval median fin with a mesenchyme core derived from LPM and propose it as an intermediate between PM-derived median fins and LPM-derived paired fins. While apparently conserved from cyclostomes to amphibia, most extant chondrichthyan embryos do not appear to have a discernible PAFF. We used micro-computed tomography (microCT) to examine prehatching stages of the elasmobranch Epaulette shark Hemiscyllium ocellatum, which did not show the presence of a clear pre-anal fin (Extended Data Fig. 9a–i). Although this pattern appears to support a loss of the PAFF somewhere in the chondrichthyan lineage, we cannot rule out the possibility that a PAFF arose independently in cyclostomes and Osteichthyes. It is interesting to note that embryos and adults of the frilled shark, Chlamydoselachus, do show bifurcated tropeic folds in the ventral midline (Extended Data Fig. 9j–m), structures previously invoked as supporting the fin fold hypothesis33.
PAFFs are mostly transient larval structures that generally lack a mineralized skeleton, which may explain their poor documentation in the fossil record. Although a true PAFF does not persist into adulthood in most species, it is seen in hagfish adults, where unlike the caudal median fin, it does not possess cartilage rays34. In the fossil record, a pre-anal fin is seen in some stem vertebrate fossils, including Haikouella and Haikouichthys, while the pre-anal fin of Kerreralepis shows well-developed plates35,36. Zebrafish larval fin mesenchyme is known to persist and contribute to the lepidotrichia of adult fins18, and LPM-derived dermal fibroblasts form cartilage during axolotl limb regeneration37. These data suggest the potential of the PAFF mesenchyme to generate skeletal outcomes.
Spontaneous duplication in twin-tail goldfish species indicates that paired LPM-derived PAFFs could have readily arisen during evolution. In Ranchu, we frequently observed multiple PAFFs, permitting retention of a median PAFF and divergence of duplicates into paired fins. Although there is no evidence of lateral fin folds present in extant vertebrates, adult specimens of the hagfish species Neomyxine biniplicata possess anterior lateral paired folds that terminate close to the pre-anal fin38, resembling partial duplications of the pre-anal fin, although their relevance to paired fin evolution is disputed39. In the fossil record, there is evidence that both mineralized and unmineralized lateral paired fin folds were common in anaspids, although homology of anapsid and gnathostome paired fins is highly contentious, as is their relevance to paired fin evolution and the lateral fin fold hypothesis1,11. Pharyngolepis, Cowielepis and Euphanerops display pre-anal ventrolateral paired ribbon structures or triangular fins with radials, while in Jamoytius, the ribbons lacked mineralized structures12,40,41. The galeaspid, Tujiaaspis, has been recently described bearing skeletal ventrolateral fins composed of skeletal units42. Subsequent regionalization of these adult paired elongate fins through anterior restriction to the pectoral fin region is proposed and exemplified by Pharyngolepis and Rhyncholepis11,43.
Our work gives weight to a model for paired fin evolution through co-option of a larval median fin programme to the LPM followed by fin duplication and subsequent regionalization to pectoral and pelvic fins. We present one possible model for how the PAFF may have led to the generation of LPM-derived paired fins (Fig. 5). The PAFF may have originated as a small ectoderm-only fin fold (as seen in amphioxus larvae44) with an LPM contribution evolving subsequently upon alterations in LPM topology, such as persistence of a somatopleure and/or a lateral mesodermal divide45,46. Similar LPM tissue contexts may have then led to PAFF elongation and following duplication, paired fin regionalization. As the PAFF possesses characteristics of both unpaired and paired fins, the PAFF may represent a novel evolutionary module or at least demonstrate components of the developmental mechanisms that contributed to the emergence of paired appendages.
Simplified evolutionary scenario of vertebrates showing the presence of a PAFF and subsequent modifications leading to paired fins. Dashed lines and dagger symbols indicate extinct lineages, and solid lines indicate extant lineages. PM-derived fins and fin folds are in cyan, while LPM-derived fins are in pink. Larval PAFF is hatched. Black arrows indicate the position of the anus.
Methods
Animal husbandry and experiments
Adult zebrafish and medaka were maintained under standard conditions, and embryos were obtained through natural crosses, kept in E3 medium at 28.5 °C and staged according to Iwamatsu47 and Kimmel et al.48. Embryos of twin-tail goldfish (Ranchu strain) were purchased from a local breeder in Singapore, cultured in E3 medium (with methylene blue) at room temperature and staged as per Li et al.49. The experimental procedures and protocols were approved by the institutional animal care and usage committees (IACUCs) at the Institute of Molecular and Cell Biology, Agency for Science, Technology and Research, Singapore (IACUC number 140924); Nanyang Technological University (IACUC number A18002); University of Zurich and the veterinary office of the Canton of Zurich; and the University of Colorado School of Medicine Anschutz Medical Campus (protocol number 979). Paddlefish embryos (P. spathula) were obtained and staged as previously described8, and experiment protocols and animal care were approved by James Madison University (IACUC number 20-1601). Lamprey embryo work was performed based on published protocols and staged per Tahara50, with experimental protocols approved by the IACUC of the California Institute of Technology (IACUC number 1436). Experiments involving X. tropicalis and X. laevis were approved by the local ethics committee at the University of Manchester and the Home Office (number PFDA14F2D). X. laevis embryos were injected with 100 pg of the -6.35drl:EGFP construct24 together with 400 pg of in vitro-transcribed membrane–mCherry messenger RNA in each blastomere at the two-cell stage. Medaka embryos were obtained from the National University of Singapore under IACUC numbers BR19-0120 and BR22-1497. Embryos of Epaulette shark were obtained from brood stock at Monash University as approved by the Monash Animal Ethics Committee under license (number 30347) and staged per Ballard et al.51.
Fish lines and MOs
The zebrafish WT strain used in this study was AB, while the following mutants were used: frftp34 (ref. 20) and hanS6 (ref. 22). The transgenic lines used were ET37, Tg(lyve1b:DsRed2)nz101, Tg(tbx16l:GAL4-VP16)sq13; Tg(UAS:Kaede)rk8, Tg(hand2:EGFP)pd24 and Tg(drl:creERT2)cz3333; Tg(hsp70l:switch)zh701 as described previously18,23,24,52. The drl:H2B-Dendra2 transgenic line was generated by Tol2-mediated transgenesis using a plasmid construct assembled through the Multisite Gateway system with LR Clonase II Plus (Life Technologies) and components from the Tol2kit53. The medaka used was the Cab strain, and the -6.35drl:EGFP transgenic line was generated by I-SceI meganuclease-mediated transgenesis54 using a plasmid that contained the -6.35drl:EGFP sequence24 flanked by two I-SceI recognition sites. The hand2 MO sequence was (5′–3′) CCTCCAACTAAACTCATGGCGACAG and injected at 500 µM. The chordin MO sequence was (5′–3′) ATCCACAGCAGCCCCTCCATCATCC and injected at 200 µM.
In situ hybridization
In situ hybridization on zebrafish, medaka and goldfish embryos was conducted as described55. Probes used were fbln1 (ref. 56), itgb3b57, bmp1a20, zebrafish hand2 (ref. 22) and medaka hand2 (ref. 58). Goldfish hand2 probe was generated by polymerase chain reaction using goldfish complementary DNA and the primer sequences (5′–3′) ACGTTTTATGGGGAGACAACC and TAATACGACTCACTATAGGGTCTTCCTTGGCGTCTGTCTT, where the T7 RNA polymerase site is underlined. The transverse section of whole-mount in situ hybridization was achieved via cryosections. Briefly, stained embryos were embedded in agar sucrose and cryoprotected in 30% sucrose. Samples were embedded in OCT medium (Tissue-Tek) and cut into 16- to 20-µm sections in a Leica cryostat (CM3050S). Paddlefish in situ hybridization was performed as previously stated, and the paddlefish hand2 probe was used59. Whole-mount chromogenic in situ hybridizations and fluorescent in situ hybridizations and sections were performed as previously described60,61 on X. tropicalis larvae using a hand2 antisense probe from the expressed sequence tag (EST) clone (TTba012k13) identified from the X. tropicalis full-length EST library62. Lamprey in situ hybridization was accomplished using HandA as a probe according to the published protocol63.
Transplantation
The hand2 cell autonomy experiments were performed by transplanting marginal cells at gastrula stages from ET37 hosts to the margin of unlabelled WT hosts as previously described64. Donors were injected with 50 pg H2B–mCherry messenger RNA either with or without 500 µM hand2 MO. Chimeras were then raised, and PAFF mesenchyme was imaged at 3 dpf.
DiI injection of medaka
CellTracker CM-DiI Dye (C7000; Invitrogen) was microinjected using filamented glass capillaries at a concentration of 1 µg µl−1 into the LPM, just posterior to Kupffer’s vesicle at the approximately four-somites stage using Eppendorf FemtoJet.
Immunostaining
Antibody staining was executed as previously described20 using either zebrafish embryos or sections. The primary antibodies, sources and dilutions used were anti-Col2A1 (II-II6B3; DSHB; 1:100), anti-eGFP (TP401; Torrey Pines; 1:500), zns-5 (AB_10013796; ZIRC; 1:200) and anti-SM22 alpha/Transgelin (ab14106; Abcam; 1:250). Secondary antibodies were all from Invitrogen and used at 1:500: Alexa 488-conjugated donkey anti-rabbit immunoglobulin G (IgG; catalogue number A-21206), Alexa 546-conjugated donkey anti-mouse IgG (catalogue number A10036), Alexa 647-conjugated donkey anti-rabbit IgG (catalogue number A-31573) and Alexa 647-conjugated donkey anti-mouse IgG (catalogue number A-31571).
Microscopy
Confocal images were taken on an LSM800, LSM880 or LSM980 Zeiss microscope. For high magnification of bright-field or Nomarski images, a Zeiss AxioImager M2 was used. A Zeiss AxioZoom V16 was utilized to capture low magnification of bright-field images. Light-sheet fluorescence microscopy was undertaken using a Zeiss Lightsheet Z.1, and sample preparation and experimental procedures were performed based on the manufacturer’s manual. Microscopy images (fluorescence and bright field) were analysed by ZEN Blue v.3.6 (Zeiss), Fiji (ImageJ v.1.52p) and IMARIS v.9.9.1 (Oxford Instruments).
High-resolution X-ray computed tomography
Shark samples previously fixed in 4% paraformaldehyde (PFA) and dehydrated for freezer storage were stained with 1% I2 dissolved in ethanol according to Metscher65. X-ray computed tomography scans were done using a Zeiss Xradia 520 Versa system. Images were generated using Avizo 3D software (ThermoFisher Scientific).
Photoconversion
To track cell lineage, Kaede and Dendra-2 photoconversion was conducted as previously described18. In brief, 24 hpf Kaede or Dendra-2 green-expressing embryos were mounted in 3% methyl cellulose or 0.5% low-melting point agarose and imaged by confocal microscopy. A selected region of interest was photoconverted to express red fluorescence using 405-nm ultra-violet laser illumination. Embryos were imaged immediately after photoconversion by sequential acquisition using 488- and 561-nm-wavelength channels to confirm successful conversion. Then, the same embryos were re-imaged at 40, 48 or 72 hpf.
Tamoxifen treatment
To induce Cre activity in Tg(drl:creERT2, hsp70l:Switch) transgenic embryos, 4-OHT was added in E3 (final concentration of 10 µM) at the 12-somite stage. Then, 4-OHT was washed off at 24 hpf, and embryos were raised until the desired stage. For specific embryonic and larval time points, embryos were heat shocked for 1 h at 37 °C to initiate the expression of eGFP (2–3 h before fixation). Subsequently, embryos were sampled, fixed and stained with 4′,6-diamidino-2-phenylindole (DAPI), and confocal imaging was undertaken.
Adult zebrafish were heat shocked for 16 h at 37 °C, anesthetized with Tricaine and euthanized in an ice-water bath. Successfully switched individuals were selected based on eGFP-expressing heart (major derivate organ of the drl-expressing LPM), and the paired pectoral and pelvic fins and anal fin were dissected and fixed overnight with 4% PFA.
Statistical analysis and reproducibility
Statistical analysis was performed using Prism v.9 (GraphPad). A two-tailed Mann–Whitney test was used when two conditions were compared. No statistical methods were used to predetermine sample size. All experiments were performed at least three times on different weeks with different biological samples. The experiments were not randomized, and investigators were not blinded. Consistent labelling within each batch of embryos was confirmed, and representative samples were used for imaging.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data Availability
The authors confirm that all relevant data are provided in this paper and in its Extended Data files. The data for measurements of fin size and cell number in Extended Data Fig. 3m–o are available in Figshare with the identifier https://doi.org/10.6084/m9.figshare.22269769. Source data are provided with this paper.
References
Coates, M. I. The evolution of paired fins. Theory Biosci. 122, 266–287 (2003).
Freitas, R., Zhang, G. & Cohn, M. J. Evidence that mechanisms of fin development evolved in the midline of early vertebrates. Nature 442, 1033–1037 (2006).
Gegenbaur, C. Elements of Comparative Anatomy (Macmillan and Company, 1878).
Balfour, F. M. On the development of the skeleton of the paired fins of Elasmobranchii, considered in relation to its bearings on the nature of the limbs of the Vertebrata. Proc. Zool. Soc. Lond. 49, 656–670 (1881).
Mivart, S. G. Notes on the fins of elasmobranchs, with considerations on the nature and homologues of vertebrate limbs. Trans. Zool. Soc. Lond. 10, 439–484 (1879).
Gillis, J. A., Dahn, R. D. & Shubin, N. H. Shared developmental mechanisms pattern the vertebrate gill arch and paired fin skeletons. Proc. Natl Acad. Sci. USA 106, 5720–5724 (2009).
Diogo, R. Cranial or postcranial—dual origin of the pectoral appendage of vertebrates combining the fin-fold and gill-arch theories? Dev. Dyn. 249, 1182–1200 (2020).
Bemis, W. E. & Grande, L. in Mesozoic Fishes 2: Systematics and Fossil Record. (eds Arratia, G. & Schultze, H.-P.) 41–68 Vrelag Dr. F. Pfeil, 1999).
Zhang, X. G. & Hou, X. G. Evidence for a single median fin-fold and tail in the Lower Cambrian vertebrate, Haikouichthys ercaicunensis. J. Evol. Biol. 17, 1162–1166 (2004).
Shu, D. G. et al. Lower Cambrian vertebrates from south China. Nature 402, 42–46 (1999).
Wilson, M. V. H., Hanke, G. F. & Märss, T. in Major Transitions in Vertebrate Evolution (eds Anderson, J. & Sues, H.-D.) Ch. 3, 122–149 (Indiana Univ. Press, 2007).
Sansom, R. S., Freedman, K. I. M., Gabbott, S. E., Aldridge, R. J. & Purnell, M. A. Taphonomy and affinity of an enigmatic Silurian vertebrate, Jamoytius kerwoodi White. Palaeontology 53, 1393–1409 (2010).
Sansom, R. S., Gabbott, S. E. & Purnell, M. A. Unusual anal fin in a Devonian jawless vertebrate reveals complex origins of paired appendages. Biol. Lett. 9, 20130002 (2013).
Lee, R. T., Thiery, J. P. & Carney, T. J. Dermal fin rays and scales derive from mesoderm, not neural crest. Curr. Biol. 23, R336–R337 (2013).
Taniguchi, Y. et al. Mesodermal origin of median fin mesenchyme and tail muscle in amphibian larvae. Sci. Rep. 5, 11428 (2015).
Parichy, D. M., Elizondo, M. R., Mills, M. G., Gordon, T. N. & Engeszer, R. E. Normal table of postembryonic zebrafish development: staging by externally visible anatomy of the living fish. Dev. Dyn. 238, 2975–3015 (2009).
Shimada, A. et al. Trunk exoskeleton in teleosts is mesodermal in origin. Nat. Commun. 4, 1639 (2013).
Lee, R. T., Knapik, E. W., Thiery, J. P. & Carney, T. J. An exclusively mesodermal origin of fin mesenchyme demonstrates that zebrafish trunk neural crest does not generate ectomesenchyme. Development 140, 2923–2932 (2013).
Okuda, K. S. et al. A zebrafish model of inflammatory lymphangiogenesis. Biol. Open 4, 1270–1280 (2015).
Asharani, P. V. et al. Attenuated BMP1 function compromises osteogenesis, leading to bone fragility in humans and zebrafish. Am. J. Hum. Genet. 90, 661–674 (2012).
Prummel, K. D. et al. Hand2 delineates mesothelium progenitors and is reactivated in mesothelioma. Nat. Commun. 13, 1677 (2022).
Yelon, D. et al. The bHLH transcription factor hand2 plays parallel roles in zebrafish heart and pectoral fin development. Development 127, 2573–2582 (2000).
Yin, C., Kikuchi, K., Hochgreb, T., Poss, K. D. & Stainier, D. Y. R. Hand2 regulates extracellular matrix remodeling essential for gut-looping morphogenesis in zebrafish. Dev. Cell 18, 973–984 (2010).
Mosimann, C. et al. Chamber identity programs drive early functional partitioning of the heart. Nat. Commun. 6, 8146 (2015).
Richardson, M. K. & Wright, G. M. Developmental transformations in a normal series of embryos of the sea lamprey Petromyzon marinus (Linnaeus). J. Morphol. 257, 348–363 (2003).
Kemp, A. Rearing of embryos and larvae of the Australian Lungfish, Neoceratodus forsteri, under laboratory conditions. Copeia 1981, 776–784 (1981).
Nieuwkoop, P. D. & Faber, J. Normal Table of Xenopus laevis (Daudin): A Systematical and Chronological Survey of the Development from the Fertilized Egg till the End of Metamorphosis 1 edn (Garland Publishing, 1994).
Hammerschmidt, M. et al. dino and mercedes, two genes regulating dorsal development in the zebrafish embryo. Development 123, 95–102 (1996).
Kon, T. et al. The genetic basis of morphological diversity in domesticated goldfish. Curr. Biol. 30, 2260–2274.e6 (2020).
Abe, G. et al. The origin of the bifurcated axial skeletal system in the twin-tail goldfish. Nat. Commun. 5, 3360 (2014).
Watase, S. On the caudal and anal fins of goldfishes. J. Sci. Coll. Imp. Univ. Tokyo 1, 247–267 (1887).
Yonei-Tamura, S. et al. Competent stripes for diverse positions of limbs/fins in gnathostome embryos. Evol. Dev. 10, 737–745 (2008).
Smith, B. G. in The Bashford Dean Memorial Volume: Archaic Fishes Vol. 2 (ed. E. W. Gudger) Article VI, 333–506 (The American Museum of Natural History, 1937)
Cole, F. J. A monograph on the general morphology of the myxinoid fishes, based on a study of Myxine Part 1. The anatomy of the skeleton. Trans. R. Soc. Edinb. 41, 749–788 (1905).
Larouche, O., Zelditch, M. L. & Cloutier, R. A critical appraisal of appendage disparity and homology in fishes. Fish Fish. 20, 1138–1175 (2019).
Blom, H. New birkeniid anaspid from the Lower Devonian of Scotland and its phylogenetic implications. Palaeontology 55, 641–652 (2012).
Kragl, M. et al. Cells keep a memory of their tissue origin during axolotl limb regeneration. Nature 460, 60–65 (2009).
Richardson, L. R. Neomyxine n.g. (Cyclostomata) based on Myxine biniplicata Richardson and Jowett 1951, and further data on the species. Trans. R. Soc. N. Z. 81, 379–383 (1953).
Donoghue, P. C., Forey, P. L. & Aldridge, R. J. Conodont affinity and chordate phylogeny. Biol. Rev. Camb. Philos. Soc. 75, 191–251 (2000).
Janvier, P. & Arsenault, M. The anatomy of Euphanerops longaevus Woodward, 1900, an anaspid-like jawless vertebrate from the Upper Devonian of Miguasha, Quebec, Canada. Geodiversitas 29, 143–216 (2007).
Blom, H. A new anaspid fish from the Middle Silurian Cowie Harbour fish bed of Stonehaven, Scotland. J. Vert. Paleontol. 28, 594–600 (2008).
Gai, Z. et al. Galeaspid anatomy and the origin of vertebrate paired appendages. Nature 609, 959–963 (2022).
Ritchie, A. The late Silurian anaspid genus Rhyncholepis from Oesel, Estonia, and Ringerike, Norway. Am. Mus. Novit. 2699, 1–18 (1980).
Mansfield, J. H. & Holland, N. D. Amphioxus tails: source and fate of larval fin rays and the metamorphic transition from an ectodermal to a predominantly mesodermal tail. Acta Zool. 96, 117–125 (2015).
Tulenko, F. J. et al. Body wall development in lamprey and a new perspective on the origin of vertebrate paired fins. Proc. Natl Acad. Sci. USA 110, 11899–11904 (2013).
Nuño de la Rosa, L., Müller, G. B. & Metscher, B. D. The lateral mesodermal divide: an epigenetic model of the origin of paired fins. Evol. Dev. 16, 38–48 (2014).
Iwamatsu, T. Stages of normal development in the medaka Oryzias latipes. Mech. Dev. 121, 605–618 (2004).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Li, I. J., Lee, S. H., Abe, G. & Ota, K. G. Embryonic and postembryonic development of the ornamental twin-tail goldfish. Dev. Dyn. 248, 251–283 (2019).
Tahara, Y. Normal stages of development in the lamprey, Lampetra reissneri (Dybowski). Zool. Sci. 5, 109–118 (1988).
Ballard, W. W., Mellinger, J. & Lechenault, H. A series of normal stages for development of Scyliorhinus canicula, the lesser spotted dogfish (Chondrichthyes: Scyliorhinidae). J. Exp. Zool. 267, 318–336 (1993).
Okuda, K. S. et al. lyve1 expression reveals novel lymphatic vessels and new mechanisms for lymphatic vessel development in zebrafish. Development 139, 2381–2391 (2012).
Kwan, K. M. et al. The Tol2kit: a multisite gateway-based construction kit for Tol2 transposon transgenesis constructs. Dev. Dyn. 236, 3088–3099 (2007).
Thermes, V. et al. I-SceI meganuclease mediates highly efficient transgenesis in fish. Mech. Dev. 118, 91–98 (2002).
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 3, 59–69 (2008).
Feitosa, N. M. et al. Hemicentin 2 and Fibulin 1 are required for epidermal-dermal junction formation and fin mesenchymal cell migration during zebrafish development. Dev. Biol. 369, 235–248 (2012).
Ablooglu, A. J., Kang, J., Handin, R. I., Traver, D. & Shattil, S. J. The zebrafish vitronectin receptor: characterization of integrin alphaV and beta3 expression patterns in early vertebrate development. Dev. Dyn. 236, 2268–2276 (2007).
Kaneko, H., Nakatani, Y., Fujimura, K. & Tanaka, M. Development of the lateral plate mesoderm in medaka Oryzias latipes and Nile tilapia Oreochromis niloticus: insight into the diversification of pelvic fin position. J. Anat. 225, 659–674 (2014).
Davis, M. C., Dahn, R. D. & Shubin, N. H. An autopodial-like pattern of Hox expression in the fins of a basal actinopterygian fish. Nature 447, 473–476 (2007).
Lea, R., Bonev, B., Dubaissi, E., Vize, P. D. & Papalopulu, N. Multicolor fluorescent in situ mRNA hybridization (FISH) on whole mounts and sections. Methods Mol. Biol. 917, 431–444 (2012).
Lea, R., Papalopulu, N., Amaya, E. & Dorey, K. Temporal and spatial expression of FGF ligands and receptors during Xenopus development. Dev. Dyn. 238, 1467–1479 (2009).
Gilchrist, M. J. et al. Defining a large set of full-length clones from a Xenopus tropicalis EST project. Dev. Biol. 271, 498–516 (2004).
Cerny, R. et al. Evidence for the prepattern/cooption model of vertebrate jaw evolution. Proc. Natl Acad. Sci. USA 107, 17262–17267 (2010).
Kemp, H. A., Carmany-Rampey, A. & Moens, C. Generating chimeric zebrafish embryos by transplantation. J. Vis. Exp. 29, 1394 (2009).
Metscher, B. D. MicroCT for comparative morphology: simple staining methods allow high-contrast 3D imaging of diverse non-mineralized animal tissues. BMC Physiol. 9, 11 (2009).
Acknowledgements
We thank K. Poss and M. Tanaka for providing zebrafish and medaka hand2 plasmids, respectively; Z. Gong for the transgenic fish line TgBAC(hand2:EGFP); D. Traver for input on the first iteration of Dendra2 transgenics; Y.-K. Tea for input; and K. P. Lim of the Lee Kong Chian Natural History Museum, Singapore for assisting with the frilled shark specimen. We also thank the Nanyang Technological University Zebrafish facility, the NTU Optical Bio-Imaging Centre (NOBIC), Colorado University (CU) Anschutz zebrafish care staff and all past and present members of our teams for continued support. This work was funded by the Industry Aligned Fund (IAF) Agency for Science, Technology and Research (grant to T.J.C. and K.-W.T.); Ministry of Education (MoE) Tier 3 (grant 2016-T3-1-005 to T.J.C., C.W. and H.M.); Ministry of Education (MoE) Tier 1 (grant 2016-T1-001-055 to T.J.C. and C.Z.); Ministry of Education (MoE) Tier 2 (grant MOE-T2EP30221-0008 to C.W.); the Company of Biologists (travelling fellowship to M.J.T.); the National Science Foundation (grants IOS-1853949 to M.C.D. and 2203311 to C.M.); the Swiss National Science Foundation Sinergia (grant CRSII5_180345 to C.M.); the Swiss Bridge Foundation (C.M.); Additional Ventures Single Ventricle Research Fund (SVRF) (grant 1048003 to C.M.); the University of Colorado School of Medicine Anschutz Medical Campus and the Children’s Hospital Colorado Foundation (C.M.); the National Institutes of Health (NIH), National Institute of General Medical Sciences (grants 1T32GM141742-01 to H.R.M. and 3T32GM121742-02S1 to H.R.M.); Australian Research Council (discovery grant DP200103219 to F.J.T. and P.D.C); National Health and Medical Research Council (senior principal research fellow APP1136567 to P.D.C.); and the NIH (grant R35NS111564 to J.S. and M.E.B.).
Author information
Authors and Affiliations
Contributions
Experiments were conceived and designed by T.J.C., C.M., K.-W.T., M.E.B., M.C.D., E.A., P.D.C., C.W. and H.H.R. and were performed and analysed by K.-W.T., R.L.L., K.D.P., J.S., H.M., H.R.M., A.N.C., Y.L., R.L., K.D., M.J.T., C.Z., E.C.B., W.T.W. and F.J.T. The manuscript was written by K.-W.T. and T.J.C., with editing and input from C.M., J.S., A.N.C., M.C.D., F.J.T., P.D.C., R.L.L., K.D.P. and E.A.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks William Bemis, Ann Burke, Brian Metscher and Oliver Stone for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 A non-paraxial mesoderm derived median fin fold.
a, Confocal image of 3 dpf Tg(tbx16l:GAL4-VP16); Tg(UAS:Kaede) embryo with paraxial mesoderm labelled by photoconverted Kaede (magenta). Unlabelled PAFF underscored by yellow bracket. c-d, Expression of Kaede in the dorsal caudal fin fold of 3 dpf Tg(tbx16l:GAL4-VP16); Tg(UAS:Kaede) is not due to localised de novo expression from the tbx16l promoter. Confocal images of 24 hpf trunks of both prior to (b) and after (c) UV photoconversion. Unconverted Kaede is in the green channel overlaid with converted Kaede in magenta. Region of paraxial mesoderm conversion and Nomarski image given in (b). At 3 dpf, converted cells can be seen in the adjacent fin fold dorsally indicating Kaede is reporting lineage (d). e, Nomarski image of PAFF at 3 dpf showing presence of mesenchymal cells. f–h, Confocal images of the pre-anal fin fold (f), dorsal (g) and caudal (h) regions of the caudal fin fold of the ET37 Enhancer Trap transgenic line at 3 dpf, indicating PAFF contains numerous mesenchymal cells which are morphologically comparable to mesenchyme of other larval fin folds. i–j, In situ hybridisation of fin mesenchyme markers itgb3b (i; 3 dpf) and bmp1a (j; 5 dpf) in both PAFFs (yellow) and caudal fin folds (cyan arrowhead). Scale Bars: 100 µm (a,f,i), 50 µm (c,d,e,j), 20 µm (g,h).
Extended Data Fig. 2 Expression of hand2 in the pre-anal fin fold.
a–b, High (a) and low (b) magnification confocal images of Tg(hand2:EGFP) zebrafish embryos at 3 dpf (a) and 8 dpf (b) showing eGFP labelling of mesenchyme of the PAFF (yellow outline, a,b), but not the caudal fin fold (cyan outline, b). c–d, In situ hybridisation of hand2 at 5 dpf showing expression of hand2 only in the PAFF (c). Higher magnification with Nomarski optics indicates expression in mesenchyme (d). Scale Bars: 100 µm (b,c), 50 µm (d), 20 µm (a).
Extended Data Fig. 3 Loss of Hand2 leads to reduction of pre-anal fin folds but not caudal fin folds.
a–l, Lateral Nomarski images of uninjected WT (a,c,e,g), hans6 mutants (b,h) and hand2 morphants (d,f) at 3 dpf (a–d) or 5 dpf (e–h). i–l, Confocal images of 3 dpf (i,j) and 5 dpf (k,l) ET37 Enhancer Trap transgenic larvae, uninjected (i,k) or injected with a hand2 morpholino (j,l). m,n Quantification of 3 dpf fin height of PAFFs (yellow) and caudal fin folds (cyan) in uninjected WT, hand2 mutants (hans6; m) and hand2 morphants (n). Loss of Hand2 leads to significant reduction of PAFFs but not caudal fin folds. n = 8 (WT), n = 15 (hans6), n = 5 (hand2 MO). Location of measurements of PAFF and caudal fin shown in (a) and (b) by yellow and cyan lines respectively. ***: p = 0.000008; **: p = 0.008; ns: p = 0.12 (m); ns: p > 0.9999 (n). o, Quantification of mesenchymal cell numbers in pre-anal (yellow bars) and ventral caudal (cyan bars) fin folds at 3 dpf. There is a significant reduction of PAFF mesenchyme in hand2 morphants compared to WT, but ventral caudal fin mesenchymal cell numbers are unaffected. n = 5 (WT), n = 12 (hand2 MO). ***: p = 0.000323; ns: p = 0.06. m–o: Data are presented as mean values with error bars representing standard deviations. In all cases n refers to biologically independent embryos. Two-sided Mann-Whitney test. Scale Bars: 100 µm (d,f,j,l), 200 µm (b,h).
Extended Data Fig. 4 Hand2 acts cell autonomously in PAFF mesenchyme.
a–d, Lateral confocal images of PAFF of 3 dpf embryos. Cells were transplanted into WT hosts from ET37 embryos that were injected with H2B-mCherry mRNA (a,b) or co-injected with H2B-mCherry mRNA and 500 µM hand2 MO (c,d). Fluorescent confocal images of mCherry (magenta) and eGFP (green) (a,c) are overlayed on Nomarski image (b,d). Arrowheads indicate transplanted cells from hand2 MO injected donors (c). Scale Bar: 20 µm (d).
Extended Data Fig. 5 Lineage tracing of the LPM into paired fins and the PAFF.
a–g, Permanently labelled LPM cells can be seen within pectoral and pelvic fins and pre-anal fin fold (arrowheads). Confocal images of the pectoral fin at 4.5 dpf (a), adult pelvic fin (b–d) and pre-anal fin fold at 5 dpf (e–g) of Tg(drl:creERT2); Tg(hsp70l:Switch) transgenics following Hydroxytamoxifen treatment and heat-shock prior to imaging. Pelvic fins and PAFF are shown as transverse sections (b–g) and were fluorescently immunostained for eGFP (b,e), Zns5 (c) or Transgelin (f). Merged images are shown in (d,g). LPM-derived mesenchyme cells of the pectoral and pre-anal fin folds are indicated by white and yellow arrowheads respectively (a, e–g). LPM-derived Zns5-positive osteoblasts of the pelvic fins are indicated by white arrowheads (b–d). Scale Bars: 20 µm (d,g), 50 µm (a).
Extended Data Fig. 6 Pre-anal fin fold mesenchyme originates from posterior-most drl-positive cells.
a–d, Lateral (a) and ventral (b–d) views of 8ss (a–b) or 10ss (c–d) drl:H2B-Dendra2 transgenic line either prior to (a–b) or following (c–d) photoconversion of posterior-most LPM (region outlined by dotted line) using UV laser illumination. e–p, Lateral confocal images of the PAFF region at 40 hpf (e–g, k–m) and 48 hpf (h–j, n–p) of a photoconverted embryo (e–j) and an unconverted control (k–p). Panels show Dendra2-red (f,i,l,o), Dendra2-green (g,j,m,p) and merged (e,h,k,n) channels. PAFFs are indicated by yellow arrowheads, and are visible in green channel in both converted and controls, but only converted embryos show red labelled PAFFs (f,i; n = 4 photoconverted fish). Magnified views of panels (f,g,i,j,l,m,o,p) are given in panels (q–x) respectively. Scale Bars: 200 µm (b), 50 µm (d,x), 100 µm (k).
Extended Data Fig. 7 LPM origin of PAFF mesenchyme is conserved in medaka, paddlefish and Xenopus.
a–c, In situ hybridisation of medaka hand2 at Stage 39 (a) and Stage 40 (b–c) showing exclusive expression in pre-anal fin mesenchyme (yellow arrowhead, b), but not in the caudal fin mesenchyme (cyan outline). Ventral caudal fin imaged in (a). d–e, Lateral confocal images of PAFF of Stage 36 transgenic Tg(-6.35drl:EGFP) medaka embryo. Fluorescent confocal images of eGFP (d) are overlayed on Nomarski image (e). Faint eGFP perdurance is seen in nascent PAFF mesenchymal cells (yellow arrowheads and inset). f–i, Ventral (f,h) and lateral (g,i) confocal images of medaka embryo injected with DiI in the posterior LPM at Stage 20 (f,h) and traced to PAFF at Stage 40 (g,i). DiI within the mesenchyme indicated by yellow arrowheads, with higher magnification example shown in inset (g,i). DiI signal (f,g) is overlayed with Nomarski (h,i). Location of paraxial mesoderm (PM), Kupffer’s vesicle (KV) and region of posterior LPM are indicated in green, red, and white respectively in (h). j–l, In situ hybridisation of hand2 in Stage 38 (j) and Stage 39 (k–l) paddlefish embryos imaged laterally (j,k) or in transverse section (l). hand2-positive PAFF fin mesenchyme indicated by yellow arrowheads is located distally. Expression of hand2 is also seen in the nascent pelvic fins at Stage 39 (green arrowheads k,l). m–o, Confocal images of a region of Xenopus laevis PAFF at NF Stage 42 injected with -6.35drl:EGFP. Lateral x-y view (n) is shown with orthogonal sections in the x-z plane (m) and y-z plane (o). Transient mosaic expression in the interstitial mesenchyme is highlighted with yellow arrowheads (m–n). Scale Bars: 20 µm (a,d), 100 µm (b,i), 50 µm (b – inset, l,o), 200 µm (h,j,k).
Extended Data Fig. 8 Reduced Chordin leads to paired duplication of PAFF.
a–b, Low (a) and high (b) power Nomarski images of duplicated PAFFs (yellow arrowheads) in 5 dpf larvae injected with chrd morpholino. c–j, Confocal images of PAFFs from 8 dpf Tg(hand2:EGFP) either uninjected (c,e) or injected with chrd morpholino (d,f,g–j). Confocal projections (c,d,i,j) and total surface renderings (e,f,g,h) highlight duplicated PAFFs (blue and yellow, f) compared to single PAFF in uninjected larvae (grey, e). Imaging of left (g) and right (h) duplicated PAFFs of chrd morphants indicated eGFP positive mesenchymal cells populated both fin folds (yellow arrowheads, i,j). k– l, Lateral (k) and ventral (l) Nomarski images of the duplicated PAFFs of the Ranchu goldfish strain at 6 dpf. Duplicated PAFFs and ventral caudal fin folds indicated by yellow and cyan arrowheads, respectively. m–n, Lateral (m) and transverse (n) views of 7 dpf Ranchu larvae stained by in situ hybridisation for hand2. Expression in individual mesenchymal cells of the PAFF indicated by yellow arrowheads (m). Occurrence of three PAFFs with core hand2 expression indicated by yellow arrowheads (n). Scale Bars: 100 µm (a,e,g), 50 µm (b), 500 µm (k), 200 µm (l), 20 µm (m,n).
Extended Data Fig. 9 Variable presence of a pre-anal fin in sharks.
a–i, The Epaulette shark does not possess a PAFF. Whole mount (a,d,g) and virtual cross sections from microCT scans (b–c, e–f, h–i) of the Epaulette shark Hemiscyllium ocellatum, at the developmental stages of 25 (a–c), 27 (d–f), and 30 (g–i). Virtual cross sections for each stage show the region either anterior (Pre-anal; b,e,h), or posterior (Post-anal; c,f,i) to the developing cloaca (white arrow, a,d,g). The dashed lines (a,d,g) indicate approximate pre-anal (magenta) and post-anal (cyan) microCT section locations (separate specimens). The developing pelvic and median fin folds are indicated by green and cyan arrowheads respectively (b–c, e–f, h–i). j–m: The tropeic folds found in a pre-anal position of the adult frilled shark, Chlamydoselachus anguineus, shown in lateral (j) and ventral (k) views (yellow arrowheads) (adult male specimen, ZRC 54430 - LKC Natural History Museum, Singapore). l–m, Ventral view of another adult frilled shark exhibits partially paired tropeic folds (yellow arrowheads) in anterior half (l), which then merges to become singular (yellow arrowhead) in posterior half (m) (anterior: left, CSIRO H 7115-01, adult male 1310 mm TL, Tasmania, Australia). Scale Bars: 1 mm (a,d,g), 5 cm (k).
Supplementary information
Supplementary Video 1
A 3D reconstruction of the duplicated PAFFs in chrd morphants. 3D rotation, surface rendering and fly-through of 8 dpf Tg(hand2:EGFP) larvae uninjected (left larva) or injected with chrd morpholino (right larva), with eGFP-positive mesenchyme visible in all PAFFs.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tzung, KW., Lalonde, R.L., Prummel, K.D. et al. A median fin derived from the lateral plate mesoderm and the origin of paired fins. Nature 618, 543–549 (2023). https://doi.org/10.1038/s41586-023-06100-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06100-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.